Template Modeling Score (bioinformatics)
The Template Modeling Score or TM-score is a measure of similarity between two protein structures with different tertiary structures. The TM-score is intended as a more accurate measure of the quality of full-length protein structures than the often used RMSD and GDT measures. The TM-score indicates the difference between two structures by a score between ![(0,1]](/2012-wikipedia_en_all_nopic_01_2012/I/668c7b55a37300c330dcd565d9e076da.png) , where 1 indicates a perfect match between two structures[1]. Generally scores below 0.20 corresponds to randomly chosen unrelated proteins whereas structures with a score higher than 0.5 assume roughly the same fold[2]. A quantitative study [3] shows that proteins of TM-score = 0.5 have a posterior probability of 37% in the same CATH Topology family and of 13% in the same SCOP Fold family. The probabilities increase rapidly when TM-score >0.5. The TM-score is designed to be independent of protein lengths.
, where 1 indicates a perfect match between two structures[1]. Generally scores below 0.20 corresponds to randomly chosen unrelated proteins whereas structures with a score higher than 0.5 assume roughly the same fold[2]. A quantitative study [3] shows that proteins of TM-score = 0.5 have a posterior probability of 37% in the same CATH Topology family and of 13% in the same SCOP Fold family. The probabilities increase rapidly when TM-score >0.5. The TM-score is designed to be independent of protein lengths.
Contents |
The equation
where 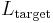 and
and 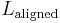 are the lengths of the target protein and the aligned region respectively.
are the lengths of the target protein and the aligned region respectively. 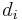 is the distance between the
is the distance between the 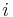 th pair of residues and
th pair of residues and
is a distance scale that normalizes distances.
See also
- RMSD — a different structure comparison measure
- GDT — a different structure comparison measure
- LCS — a different structure comparison measure
References
- ^ Zhang Y and Skolnick J (2004). "Scoring function for automated assessment of protein structure template quality". Proteins 57 (4): 702–710. doi:10.1002/prot.20264. PMID 15476259.
- ^ Zhang Y and Skolnick J (2005). "TM-align: a protein structure alignment algorithm based on the TM-score". Nucleic Acids Res 33 (7): 2302–2309. doi:10.1093/nar/gki524. PMC 1084323. PMID 15849316. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1084323.
- ^ Xu J and Zhang Y (2010). "How significant is a protein structure similarity with TM-score=0.5?". Bioinformatics 26 (7): 889–895. doi:10.1093/bioinformatics/btq066. PMC 2913670. PMID 20164152. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2913670.
External links
- TM-score webserver — by the Yang Zhang research group. Calculates TM-score and supplies source code.
![\text{TM-score}=\max\left[ \frac{1}{L_\text{target}}\sum_i^{L_\text{aligned}}\frac{1}{1%2B\left(\frac{d_i}{d_0(L_\text{target})}\right)^2} \right]](/2012-wikipedia_en_all_nopic_01_2012/I/d57d2f6433f00c200adebea5de4ae1b0.png)
![d_0(L_\text{target})=1.24\sqrt[3]{L_\text{target}-15}-1.8](/2012-wikipedia_en_all_nopic_01_2012/I/3fd378a62a18fdb693578b40efee813a.png)